|
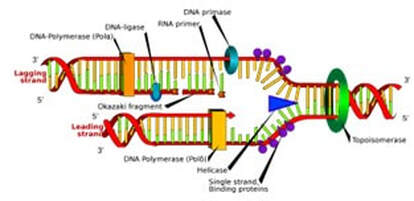
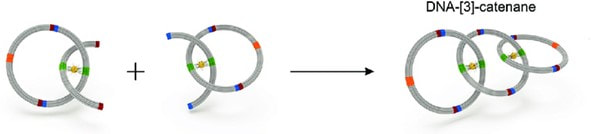
By Lucía Achucarro Rosalind Franklin is undoubtedly one of the biggest names in science. Most remembered for her discovery of DNA’s double helix structure, her multi-disciplinary approach to research is one to emulate. Franklin combined her love for chemistry with complex mathematical analyses and used X-ray crystallography as an experimental tool to provide the missing piece of the DNA puzzle. Her findings forever changed our understanding of how life works. Now, 70 years later, Dr. María José Fernández is continuing Franklin’s legacy by embracing a similar multidisciplinary approach to her research in DNA topology – the study of how DNA strands are intertwined. Fernández has modernized Franklin’s discoveries by combining mathematical analyses and coding to further advance the field. Dr. María José Fernández (Credit: Matías Nicolás del Barco Portillo, Marketing Department of the Communication Department of the FP-UNA) Born and raised in Asunción, Paraguay, Fernández describes herself as a curious kid, always wanting to know how things worked and the mechanisms behind them. Fernández didn’t always know if she wanted to be a scientist, all she knew was that she wanted a challenge. “People would ask me ‘What do you want to study?’ and I would always respond, ‘I don’t know, but it better be something difficult,’” she recounted. “I’ve always wanted to leave a mark and make a difference, in whatever field I ended up in.” Fernández graduated with a Bachelor’s Degree in Biology from the National University of Asunción in the year 2000 and secured a Gold Medal -- the equivalent of being Summa Cum Laude. She continued her academic path at the Complutense University of Madrid where she earned both her master’s and doctorate degrees in biochemistry and molecular biology. In 2009, she completed a postdoctoral fellowship program at the University of Oxford in the Weatherall Institute of Molecular Medicine. At present, she is a full-time Research Professor and the Bioinformatics Group Coordinator at the National University of Asunción. By merely looking at her academic accomplishments, we can already get a sense of Fernández’s drive and resiliency. “I’m never satisfied with my work, I always want more,” she says. “My main motivation is to generate as much knowledge as I can.” It’s safe to say that her words reflect the thinking of a true scientist — someone who, rather than idly waits for a solution, strives to be the one who comes up with it. Dr. María José Fernández (Credit: Matías Nicolás del Barco Portillo, Marketing Department of the Communication Department of the FP-UNA) This way of thinking has proven successful for Dr. Fernandez, who conducts and publishes research in several different areas like molecular pathology and cellular differentiation and cancer. Although her insight in each of these areas is incredibly fascinating and important, I’d like to focus on her work on DNA topology, the study of the DNA helix shape and path, through the use of mathematical simulations in 3D. Why study DNA Topology? In a living cell, DNA helices change their shape when they undergo replication, transcription, recombination, and repair — all very important processes that happen for organisms to function. In order to change shape accurately and effectively, certain enzymes, called topoisomerases, help regulate the movement of DNA. These enzymes introduce temporary strand breaks in the DNA and facilitate protein interactions to prevent over- or under- twisting of DNA and thus act as stress busters; without topoisomerases, the stress built on DNA due to excessive twisting can indeed hinder the aforementioned processes. Try to imagine DNA as a kind of rope with its ends tied together. As the rope twists over itself, it creates more tension, and it may create tangles or knots that, in the case of DNA, may prevent it from carrying out important life processes. In the rope scenario, topoisomerases are seen as scissors, cutting the rope at certain parts and creating transient breaks that allow the tension to dissipate and the processes to continue smoothly. However, exactly how topoisomerases regulate DNA movements is still unclear. Inhibiting topoisomerases can lead to cell death, and the ability to turn off topoisomerase activity can be very beneficial for the treatment of diseases like cancer. Indeed, most chemotherapy treatments contain agents that inhibit topoisomerases. This is one of the many reasons why research in the field of DNA Topology is essential for understanding the “Code of Life.” There’s also beneficial advantages of studying DNA replication and the work of topoisomerases in bacteria. Topoisomerases cause mutations in some bacterial strains that lead to antibiotic resistance, which can be harmful for learning about how antibiotic resistance affects us humans. If researchers can study the replication process step by step along with the topoisomerase influence in it, they may be able to pinpoint where the resistance was created and help overcome it by developing stronger antibiotics. The approach: Study DNA replication in vivo Dr. Fernandez, along with a team of researchers, chose to study DNA replication in depth to help elucidate some of the unknowns around this important process. One of their goals is to develop a new tool by which scientists can study DNA replication in its different stages in vivo. This means that they will be able to see the replication actively happening in each of its stages in a living organism — most likely a bacteria. An advantage of this effort is that by seeing replication in action, they can also see the work that topoisomerases do. Even though tools developed for similar purposes already exist (1), they contain some limitations which Fernandez and her team hope to overcome. In their journey towards developing this tool, the first step was to understand DNA replication and its movements down to its core. Part of this was documented in their revolutionary 2020 paper "Distribution of torsional stress between the un-replicated and replicated regions in partially replicated molecules" (Martínez et al., 2020) (2). This paper earned its authors a National Science Award in Paraguay (the most important award in research and development that the Paraguayan Government grants to scientists who have contributed to the gain of national and groundbreaking knowledge). The main goal of this particular study was to analyze and understand the interchange of the over-or under-winding of a DNA strand (supercoiling) and the first stage of the process by which two DNA strands link together (pre-catenation) during the replication of bacterial DNA molecules. Now, these are terms the majority of people outside the scientific world haven’t heard before, and if you’re one of them (or need a refresher) don’t worry — I’ll explain what it’s all about. What is DNA supercoiling? Dr. Fernández and her team studied a specific type of DNA: right-handed DNA double helix (also called B form DNA) molecules. The DNA molecule has four bases: adenine, cytosine, guanine, and thymine — the bases along each DNA strand pair together to form the helix. Adenine always pairs up with thymine, and guanine with cytosine. When in a relaxed state, the two DNA strands twist around their axis every 10.4 to 10.5 bases. Some enzymes have the ability to add or subtract twists in the DNA, but while doing so they impose strain. If the DNA is in a circular shape (for example, a plasmid, a small DNA molecule found in bacteria), or if the ends are tightly held forming a loop, adding or subtracting twists leads to straining, which leads to the DNA being in a so-called super-coiled state (3). Supercoiling helps accommodate the extra or missing twists by allowing the DNA to twist around itself. It can also help DNA compact inside the cell and fit in the tight space it’s given. There are two types of supercoiling: positive, when there is overtwisting, symbolized by the + sign, and negative, when there’s under twisting, symbolized by the - sign. Now that we’ve got supercoiling out of the way, let’s move on to how it affects DNA replication and where topoisomerases come into the picture. Graphic of DNA Replication (Credit: “LadyofHats” Mariana Ruiz, Public Domain, https://commons.wikimedia.org/w/index.php?curid=6916971. Torsional stress and topoisomerases Most DNA molecules are in a (-) supercoiled state, undertwisted, because this state is easier for the DNA to maintain. For example, in bacterial cells, the (-) supercoiled state makes it easier for the DNA strands to separate and advance during replication. However, DNA is forced to supercoil positively (+) during replication when DNA helicases unzip the DNA double helix. As an analogy, one can try unwinding a twisted rope by pulling the two strands apart slowly while holding the other end of the rope in place and observing overwinding of the twisted region; you can see the tension build up at the point where the twisted and untwisted region meet. Similarly, in the DNA helix, the torsional stress (the stress that acts on the DNA’s replication fork) caused by the (+) supercoiled state begins to accumulate directly in front of the replication fork (the point at which the two strands of DNA are separated to allow replication of each strand, Fig.1). This stress accumulation, in turn, prevents progression of the replication fork through the helix. This is when the topoisomerases come in and help relieve that torsional stress so that the replication process can continue smoothly. Not a lot is known regarding topoisomerases and their specific role in replication, but, researchers have come up with viable theories. One theory (4) suggests that the topoisomerases act simultaneously in the replicated and un-replicated regions of the DNA after they (+) supercoil. This diffusion can only happen by the swiveling (turning around a point or axis) of the replication fork, and through the swiveling and diffusion, the un-replicated region always stays (-) supercoiled, facilitating replication. Although the (+) supercoiling disappears by the action of topoisomerases, the resulting replicated molecule comes out heavily catenated. When a molecule is catenated, it basically means that it’s connected together like chain links. But for a catenane to be produced, pre-catenanes need to exist beforehand. In simple terms, pre-catenanes are two or more intertwined molecules that form a catenane following ring closure as shown below. Basically, what this theory suggests, is that pre-catenanes are produced during the replication process. In some way, it appears that the (+) supercoiling is swapped for pre-catenanes. So far, this topoisomerase “pathway” seems favorable as topoisomerases are able to relieve torsional stress in both regions, and the swiveling of the fork allows for easier replication But, what still remains a mystery is how the pre-catenanes are produced, whether forks are able to rotate in the first place during replication, and if so, what forces cause them to rotate. A simple example of a catenane being produced (CC BY 4.0) Model results To analyze the aforementioned pathway in action — the fork swiveling — and to understand the interchange of supercoiling and pre-catenation on DNA molecules during replication, Dr. Fernandez’s team first needed to establish what they wanted to replicate in their computer simulations. They agreed on recreating a partially replicated DNA plasmid, which is a circular molecule of DNA extracted from bacteria (they are very useful for experiments seeing as they can replicate independently of the main bacterial chromosome), in deproteinized conditions and in vitro — meaning outside of a living organism. These conditions allow them to study the fork swiveling in action without hindrance from outside factors, something that would be impossible to achieve in the presence of proteins. The plan they came up with was to use the Metropolis Montecarlo Method and a worm-like chain model (a model used to characterize the behavior of a semi-flexible polymer) to accurately depict a DNA molecule and simulate its movements and topological complexity during DNA replication. They created partially replicated circular plasmids of 4400 base pairs where either 25, 50, or 75% of the molecule was replicated in order to study the dynamics of the structural changes at different stages. The plasmids were constructed as a chain of consecutive cylinders of constant length and thickness (as seen in Fig. 2), and they programmed it so that it would move just as a real DNA molecule would during replication. Next, they worked on the process of simulating replication. Replication begins with unzipping the DNA molecule to form a replication fork, and the advance of the forks generates an over-winding of the DNA (supercoiling) that accumulates only immediately ahead of the forks, in the un-replicated region. The DNA molecule also needs to stay in thermodynamic equilibrium (a state where all the factors, like temperature and pressure, are balanced and remain the same until another external factor changes that stability) in order for the replication process to continue. Thermodynamic equilibrium is achieved when the net value of the torque (the force that causes torsional stress) between the replicated and un-replicated regions is zero. When the un-replicated region winds or unwinds, it applies torque to the fork, the same thing that happens when the replicated region braids or un-braids, but the torque is applied in the opposite direction. Because unzipping the DNA duplex results in torsional stress accumulation ahead of the fork, the torsional stress becomes uneven between the replicated and un-replicated regions. When the net torque is nonzero, the equilibrium is disturbed. Therefore, torsional stress balance between the replicated and un-replicated regions is the key to achieving net-zero torque and thus mechanical and thermodynamic equilibrium. A direct measure of torsional stress is the potential elastic energy. Hence in the simulations, the team measured the elastic energy of the replicated and un-replicated regions for 25, 50 and 75% partially replicated molecules. As expected, they found that the elastic energy in the un-replicated region is higher than in the replicated region, and this imbalance induces fork rotation. Consequently, it allows the redistribution of the torsional stress between both regions by facilitating the interconversion of the supercoiling from the un-replicated region and the precatenanes to the replicated one. At a certain point in the process, the elastic energy is balanced in both regions, which means that the net torque is zero and the DNA molecule has attained mechanical and thermodynamic equilibrium. Once this happens, the fork rotation stops and the replication process continues. This video published along with the paper showcases one of their simulations, specifically one that involves a 50% replicated plasmid. It displays how the dynamic change in the conformation of the un-replicated and replicated regions results in the redistribution of the torsional stress initially accumulated in the un-replicated one. Even if the authors' process does have some limitations — as one cannot replicate such a complex molecule exactly — the simulation procedure was able to accurately show the dynamic and equilibrium properties of supercoiled and pre-catenated DNA. Through running the simulations in 25, 50, and 75% replicated DNA plasmids, they observed the following:
Creative collaboration This was the first study of the interchange of torsional strain between the un-replicated and replicated regions in replicating DNA to use a numerical approach. Thanks to these observations, the researchers were able to find enough plausible evidence to propose that the distribution of torsional stress between the un-replicated and replicated regions drives fork swiveling, thereby allowing the interchange of supercoiling and pre-catenation in partially replicated DNA molecules. This study perfectly exemplified the ways in which biology can benefit from interdisciplinary research and collaboration. In this case, computer coding and simulation have acted not just as an extra set of tools but were the key to success. Without knowledge in this extra discipline, the researchers probably wouldn’t have been able to carry on and analyze their study as well as they did. As Dr. Fernández explained to me, her team carried out all the numerical simulations and coding. Most of the coding work was developed by Víctor Martínez with help of Hernando Maldonado, an engineering student, and Christian Schearer, a mathematician from the National University for the mathematical model. For the theoretical model and experimental work, they collaborated with Jorge B. Schvartzman and his group at the CIB-CSIC in Madrid, Spain, whose mentorship left an incredibly big mark on Dr. Fernández, especially since Dr. Schvartzman is also a Paraguayan scientist and one of the people that motivated her to enter the world of DNA Topology. In her own words, “All of our achievements were possible thanks to Jorge B. Schvartzman, the training he has given us, his passion for science and the importance he passed on to us of knowing how to communicate scientific results.” Fernández also mentioned that the whole process was an educational one, as they only had basic coding knowledge at the beginning of the study. They learned a lot from other researchers’ advice at international DNA Topology meetings, where they would present advances in their work. This only highlights the importance of scientific collaboration, as it boosts breakthroughs, especially in this particular case, as the researchers were treading through unexplored grounds and concepts. “The main challenge we had to face,” commented Fernández, “was the simulation of partially replicated DNA molecules, simply because no one had done it before.” Together, the team relied on creativity to rise to the challenge. This simplified summary of the study just merely conveys the beautiful complexity of what Fernández and her fellow researchers have done. As the years advance, more and more disciplines are finding common ground, especially with the aid of technology. And it is scientists, like Fernández, who create collaborative opportunities and trailblaze their way to the forefront of research. It’s just a simple formula of success, really. We’ve seen it work with Rosalind and we will see it work again, undoubtedly, with Dr. María José Fernández. Sources
|
Archives
February 2023
DiscovHER BlogScientista DiscovHER is a blog dedicated to discovHERies made by women in science. Follow us for links to the latest resHERch! Categories
All Alexandra Brumberg Amy Chan Avneet Soin Chemistry Diana Crow Engineering Health/medicine Indulekha Karunakaran Iqra Naveed Johanna Weker Lidiya Angelova Michael Clausen Mind Brain And Behavior Muhammad Hamza Waseem Nikarika Vattikonda Opinion Prishita Maheshwari-Aplin Technology Uma Chandrasekaran |
The Network for Pre-Professional Women in Science and Engineering
The Scientista Foundation is a registered 501(c)(3) -- Donate!




 RSS Feed
RSS Feed